Donor-Derived Myelodysplastic Neoplasm in a Pediatric Patient with Aplastic Anemia: A Case of Post-Transplant Clonal Evolution
Kiran Ghodke1,*, Nirali Sanghvi2, Sneha Ann Oommen3 and Santanu Sen4
1Consultant Hematopathologist, Hematology Department, Kokilaben Dhirubhai Ambani Hospital and Research Institute, Rao Saheb Achutrao Patwardhan Marg, Four Bungalows, Andheri (W), Mumbai, Maharashtra, India
2Senior Associate Consultant, Paediatric Hematology-Oncology and BMT unit, Kokilaben Dhirubhai Ambani Hospital and Research Institute, Rao Saheb Achutrao Patwardhan Marg, Four Bungalows, Andheri (W), Mumbai, Maharashtra, India
3Associate consultant Hematopathologist, Hematology Department, Kokilaben Dhirubhai Ambani Hospital and Research Institute, Rao Saheb Achutrao Patwardhan Marg, Four Bungalows, Andheri (W), Mumbai, Maharashtra, India
4Consultant, Paediatric Hematology-Oncology and BMT unit, Kokilaben Dhirubhai Ambani Hospital and Research Institute, Rao Saheb Achutrao Patwardhan Marg, Four Bungalows, Andheri (W), Mumbai, Maharashtra, India
Received Date: 28/01/2026; Published Date: 23/03/2026
*Corresponding author: Dr. Kiran Ghodke, Consultant Hematopathologist, Kokilaben Dhirubhai Ambani Hospital, Mumbai, India
Abstract
We report a pediatric case of severe Aplastic Anemia (AA) evolving into donor-derived myelodysplastic neoplasm (MDS) three years after haploidentical bone marrow transplant. The patient’s initial workup revealed hypocellular marrow without dysplasia, negative inherited-failure studies, and a heterozygous GFI1B c.503G>T variant of uncertain significance. After failing eltrombopag and ATG therapy, he underwent paternal haploidentical transplant. He engrafted successfully but remained immunocompromised, with recurrent infections. Approximately 3 years post-transplant, he developed new bicytopenia. Bone marrow evaluation showed hypercellular marrow with trilineage dysplasia and increased blasts (8–10%), consistent with MDS. Flow cytometry highlighted aberrant CD34+/CD117+ progenitors lacking CD13, and granulocytic maturation abnormalities (“myelocyte bulge (abnormal accumulation of intermediate granulocytic precursors)”), all supporting a clonal myeloid process. Targeted sequencing of the marrow revealed acquired DNMT3A (two distinct missense mutations, VAF ~33%) and BCOR frameshift mutations, not present at baseline. Donor cells were DNMT3A wild-type, confirming these mutations arose post-transplant. We discuss how this represents post-transplant clonal evolution: a donor-derived myeloid neoplasm driven by somatic epigenetic mutations. This case highlights diagnostic challenges in differentiating AA from emerging MDS, the role of flow and molecular studies in characterization, and the clinical implications of GFI1B and DNMT3A aberrations in hematopoietic clonal evolution.
Keywords: Aplastic anemia; Myelodysplastic neoplasm; Haploidentical bone marrow transplant; GFI1B; DNMT3A and BCOR
Introduction
Severe Aplastic Anemia (AA) is an immune-mediated marrow failure characterized by pancytopenia and hypocellular marrow without dysplasia [1]. Up to ~15% of AA patients can later develop myelodysplastic neoplasm (MDS) or acute leukemia, especially those unresponsive to immunosuppression [1]. Recent genomic studies have revealed that a substantial subset of AA patients harbor somatic mutations in “MDS genes” even at diagnosis – notably ASXL1, DNMT3A, TET2, and BCOR – which predict higher risk of progression to MDS/AML [1,2]. In particular, DNMT3A and ASXL1 mutations in AA are associated with clonal hematopoiesis and poorer outcomes [2,3]. Pediatric MDS is biologically distinct from adult MDS: children often have hypocellular marrows and different genetic drivers [4]. Notably, DNMT3A mutations are rare in primary pediatric MDS [4]. GFI1B is a transcription factor required for erythroid/megakaryocytic differentiation; germline GFI1B mutations cause thrombocytopenia and bleeding disorders [5]. Dominant-negative GFI1B mutations have been reported in familial syndromes predisposing to MDS/AML [5]. While the role of our patient’s GFI1B variant is unclear, it raised suspicion of an underlying predisposition to dyspoiesis. This report emphasizes the diagnostic reasoning integrating morphology, flow cytometry, and genetics, and reviews similar cases of post-transplant clonal myeloid neoplasms in the context of GFI1B and DNMT3A alterations.
Case Report
The patient, a previously healthy young boy, presented with severe pancytopenia and was diagnosed with very severe AA. Initial bone marrow biopsy in year 2022 showed a hypocellular marrow without dysplasia or abnormal clones. Laboratory workup was negative for paroxysmal nocturnal hemoglobinuria, chromosomal breakage, and viral causes. A germline panel for bone marrow failure revealed a heterozygous GFI1B c.503G>T variant (autosomal dominant model), deemed of uncertain significance and linked in literature to platelet production defects [5]. As no non-hematopoietic tissue was available, germline status was inferred. Despite immunosuppression (horse ATG, cyclosporine) and eltrombopag, blood counts did not recover. Given transfusion dependence and lack of response, the patient underwent haploidentical stem cell transplant from his father (August, 2022) using a conditioning regimen of ATG–fludarabine–cyclophosphamide–TBI–PTCy. Donor-specific antibodies were negative. He received a CD34+ cell dose of 7.2×10^6/kg. Engraftment occurred at Day+15 (WBC) and Day+21 (platelets), and full donor chimerism (>98%) was documented. The immediate post-transplant course was notable for bacterial sepsis and CMV reactivation, both treated successfully. Subsequent chimerism remained ≥95%, indicating sustained donor hematopoiesis.
Approximately 2 years post-transplant the child had recurrent Mycoplasma pneumoniae infections, treated with doxycycline. At nearly 3 years post-transplant (Sept 2025), he developed symptomatic cytopenias. Physical exam was unremarkable. Laboratory data revealed severe bicytopenia (Hb 4.6 g/dL, ANC <500/µL, platelets 73×10^3/µL) with macrocytosis (MCV 106 fL). Reticulocyte count was inappropriately low. Workup for hemolysis, micronutrient deficiency (folate slightly low, B12 normal), and infection (parvo IgM negative, viral PCR panel negative) was unrevealing.
Bone marrow aspiration and biopsy (September, 2025) demonstrated a hypercellular marrow with left-shifted but arrested myeloid maturation and multilineage dysplasia. Blasts comprised 8% of marrow cells. Megakaryocytes were decreased and dysplastic. These findings indicated a myelodysplastic/myeloid neoplasm. Immunohistochemistry (IHC) showed ~8–10% CD34+ and CD117+ blasts [Figure 1]. Flow cytometry identified an abnormal blast population (0.47% of all events) with moderate CD34 and CD117, moderate CD33, bright HLA-DR, dim to negative CD38 and uniformly negative lineage markers (CD13, CD14, CD15, CD36) [Figure 2]. Concurrently, analysis of the maturing myeloid compartment revealed aberrant granulocyte maturation: a relative increase in myelocytes (“myelocyte bulge (abnormal accumulation of intermediate granulocytic precursors)”) and asynchronous expression of CD11b, CD13, and CD16, consistent with dysplasia. Myeloid dysplasia on flow is characterized by such abnormal marker patterns [6].
The overall picture (cytopenias, dysplasia, blast increase) was diagnostic of MDS with increased blasts. The patient received transfusional support and doxycycline for a concurrent positive Mycoplasma IgM, but counts remained poor (chronic transfusion dependency). A second marrow (October, 2025) again showed dysplasia and 4% blasts by aspirate and CD34 IHC ~5–6% of blasts on trephine. Flow cytometry now detected ~2.3% aberrant myeloid blasts (CD34+/CD117+, HLA-DR+, dim to negative CD38 and negative for mature myeloid antigens), confirming persistent MDS. Conventional cytogenetics and MDS FISH panels were normal. Lineage-specific chimerism was performed using STR markers on flow-sorted CD15+ CD45+ myeloid cells. The findings confirmed 100% donor origin in this lineage. Next-generation sequencing of marrow cells identified two somatic missense DNMT3A mutations (c.2645G>A and c.2053G>A, both VAF ~33%) and a BCOR frameshift (VAF ~7%). These were absent in archived pre-transplant samples. Sanger sequencing of the donor’s blood was negative for DNMT3A mutations. Next-generation sequencing was performed on donor peripheral blood had a sensitivity of 5% variant allele frequency (VAF) for mutation detection was also negative for any other mutations. Together, the data support a post-transplant clonal evolution to MDS.
Table 1: Temporal acquisition of mutations and donor origin.

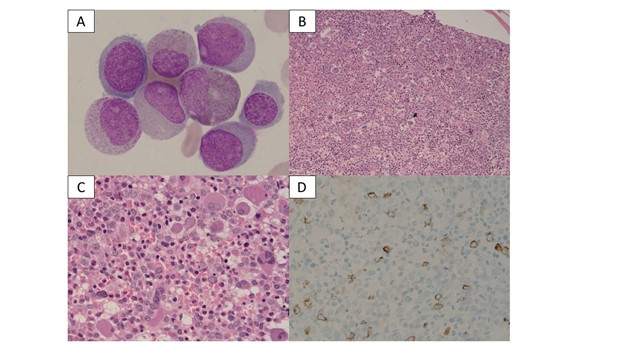
Figure 1[A-D]: Bone Marrow Morphology & Histology.
A – Bone Marrow Aspirate (100x, Wright-Giemsa): Showing an increase in blasts and a left-shifted myeloid maturation sequence accompanied by significant dyspoiesis. B – Trephine Biopsy (10x): Showing a hypercellular bone marrow with marked myeloid preponderance, left-shifted maturation, and an increase in megakaryocytes exhibiting dyspoiesis. C – Trephine Biopsy (40x): High-power view confirming myeloid preponderance with an interstitial increase in blasts. Megakaryocytes show dyspoiesis and abnormal clustering. D – Immunohistochemistry (CD34): Staining highlights the interstitial increase in CD34+ blasts and immature precursors.
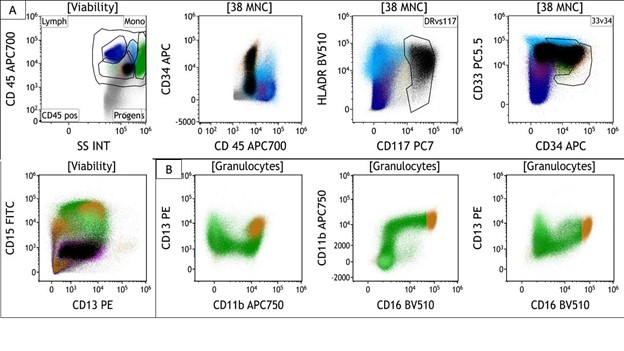
Figure 2 [A-B]: Flow Cytometric Immunophenotyping
A – Sequential Gating: Analysis of CD45+ cells and CD38 gating of mononuclear cells identifies an abnormal myeloid blast population (black). These blasts express: Positive: Heterogeneous CD34, moderate CD117, moderate CD33, and moderate HLA-DR. Negative: Uniformly negative for CD15 and other lineage-specific markers (not shown in figure). B – Myeloid Maturation Analysis: Examination of the maturing myeloid compartment reveals aberrant granulocyte maturation. Key findings include a “myelocyte bulge” (abnormal accumulation of intermediate granulocytic precursors) and asynchronous pattern expression of CD11b, CD13, and CD16.
Discussion
This case illustrates post-transplant clonal myeloid evolution in a child with prior AA. The diagnostic journey involved integrating clinical, morphologic, flow cytometric, and genetic data (Table 1). Initially, distinguishing AA from hypocellular MDS can be difficult; in AA the marrow is typically empty without dysplasia, whereas hypoplastic MDS has subtle dysplasia and genetic anomalies [1]. In our patient’s first biopsy, no dysplasia or clonal cytogenetics were seen, supporting true AA.However, years later the reappearance of dysplasia and blasts (8–10%) signaled malignant transformation. Flow cytometry was crucial to characterize the emergent clone. The blasts were CD34+ and CD117+ myeloblasts lacking mature markers (CD13, CD14, CD15), with bright HLA-DR and reduced CD38 — an immunophenotype typical for myelodysplastic blasts [6]. Granulocyte flow analysis revealed a “myelocyte bulge”: accumulation of intermediate myeloid precursors relative to mature neutrophils on CD11b/CD13/CD16 plots, a recognized immunophenotypic feature of MDS-related dysgranulopoiesis. This clustering of cells is abnormal and is seen in MDS. The combined morphologic and flow findings met WHO criteria for MDS with increased blasts [7].
The genetic findings define the clone driving this neoplasm. Two somatic DNMT3A mutations and a BCOR loss-of-function mutation were discovered in marrow cells in late 2025. Notably, these mutations were not present before transplant, and the donor was wild-type, indicating that they emerged in donor-derived hematopoietic cells post-transplant. DNMT3A and BCOR are well-known recurrent MDS/AML genes. In adult MDS, DNMT3A mutations are common and portend poor prognosis. Yeung et al. showed that DNMT3A (and ASXL1/TP53) mutations in AA significantly accelerate evolution to MDS/AML and shorten survival [3]. In AA cohorts, DNMT3A mutations often persist or expand over time [3]. BCOR mutations, while rare (<2%) in pediatric myeloid malignancies [8], are associated with high-risk MDS features: severe thrombocytopenia, increased blast burden, rapid AML transformation, and dismal outcomes [8]. The coexistence of DNMT3A and BCOR mutations in our patient’s marrow likely drove aggressive clonal expansion and marrow failure.
The detection of a GFI1B variant at initial presentation raises interesting questions. GFI1B is critical for erythroid/megakaryocytic differentiation; germline GFI1B mutations cause autosomal dominant thrombocytopenia and platelet dysfunction [5]. Recently, a dominant-negative GFI1B mutation was reported to cause familial platelet disorder progressing to AML [5]. Population databases (e.g., gnomAD) for patient’s variant (c.503G>T) showed to be ultra-rare with no known pathogenic classification. Based on ACMG criteria the variant remains of uncertain significance. It may represent a germline predisposition to marrow dysfunction, but there is no evidence it contributed to the later myeloid clone (no known myeloid mutations in GFI1B itself were found). We consider it a possible predisposing factor for his marrow failure and dysmegakaryopoiesis, but the transformation to MDS was clearly driven by acquired somatic mutations.
Importantly, this case represents a donor-derived myeloid neoplasm. After allo-transplant, a small fraction of recipients (<5%) can develop donor-cell leukemia or MDS (DCL) [9]. Clonal evolution in donor cells is rare but often carries adverse prognosis [9,10]. Our patient’s early high donor chimerism (95–98%) and paternal origin of cells with acquired mutations indicates that the MDS arose within engrafted donor hematopoiesis. This contrasts with residual host disease; our donor was tested and lacked the DNMT3A lesions. The latent interval (~36 months) is consistent with reported cases of donor-cell leukemogenesis, which can range from months to years’ post-transplant [10]. Clinically, distinguishing donor- vs. host-derived MDS is critical, as it informs donor selection for possible second transplant and prognosis [10]. In practice, this may require chimerism and molecular studies; our work-up confirmed the somatic origin.
Secondary MDS after AA, whether host- or donor-derived, is not unprecedented. AA patients who fail initial therapy have a 10–20% risk of clonal evolution [1]. This typically manifests as new cytogenetic anomalies (e.g., monosomy 7) or mutations. Our case is unusual because the new clonal mutations appeared in donor cells. Only sporadic case reports exist of donor-derived MDS from CHIP-positive donors or spontaneous mutations after transplant [9,10]. The phenomenon underscores the inherent genomic instability and selective pressures in bone marrow failure states. In our patient, chronic inflammation or infection (e.g., repeated Mycoplasma infections) might have provided stress that favored expansion of an HSC with DNMT3A/BCOR hits, although this is speculative.
From a management viewpoint, the patient remains supportive (transfusions, infection prophylaxis). The long-term outlook is concerning given high-risk molecular features. Allogeneic transplant from an alternative donor would typically be the only curative option, but the choice of donor is complex if the original donor graft itself spawned the malignancy. This underscores the dilemma in donor selection after donor-derived neoplasia: a new, unrelated donor might be preferable to avoid repeating the same clone expansion, as noted by Ihlow et al [10].
Conclusion
In summary, we describe a novel case of pediatric AA progressing to donor-derived MDS characterized by acquired DNMT3A and BCOR mutations. This case highlights several key lessons. First, persistent cytopenias in a post-transplant AA patient warrant urgent re-evaluation for clonal evolution, using morphology and flow to detect emerging dysplasia. Second, comprehensive genetic testing (including NGS panels covering MDS genes) is essential in AA and post-AA patients, as somatic mutations can signal impending transformation [2]. Third, GFI1B variants of uncertain significance may exist in AA workups; while their role remains to be fully understood, GFI1B’s function in megakaryopoiesis suggests careful monitoring of platelet lineage. Finally, post-transplant myeloid neoplasms can be of donor origin [9][10], and their recognition has implications for prognosis and future therapy. This patient’s clinical course exemplifies clonal evolution in a post-transplant setting and the need for vigilance in long-term follow-up.
Acknowledgement: We acknowledge Ms. Akshita Rane, MSc, Research Scientist, Hematology Department, Kokilaben Dhirubhai Ambani Hospital and Research Institute, Mumbai, for technical assistance with sample processing.
References
- Kulasekararaj AG, Jiang J, Smith AE, Mohamedali AM, Mian S, Gandhi S, et al. Somatic mutations identify a subgroup of aplastic anemia patients who progress to myelodysplastic syndrome. Blood, 2014; 124(17): 2698-2704. doi: 10.1182/blood-2014-05-574889.
- Wang P, Jiang W, Lai T, Liu Q, Shen Y, Ye B, et al. Germline variants in acquired aplastic anemia: current knowledge and future perspectives. Haematologica, 2024; 109(9): 2778-2789. doi: 10.3324/haematol.2023.284312.
- Ye L, Zhang L, Zhang D, Zhao X, Li Y, Xiong Y, et al. Somatic mutations and the efficacy of immunosuppressive therapy in severe/very severe aplastic anemia and transfusion-dependent nonsevere aplastic anemia. Ann Hematol, 2025; 104(8): 3981-3992. doi: 10.1007/s00277-025-06393-z.
- Kotmayer L, Kennedy AL, Wlodarski MW. Germline and somatic genetic landscape of pediatric myelodysplastic syndromes. Haematologica, 2025; 110(9): 1974-1986. doi: 10.3324/haematol.2024.285700.
- Anguita E, Candel FJ, Chaparro A, Roldán-Etcheverry JJ. Transcription Factor GFI1B in Health and Disease. Front Oncol. 2017 Mar 28;7:54. doi: 10.3389/fonc.2017.00054. PMID: 28401061; PMCID: PMC5368270.
- Bento LC, Correia RP, Pitangueiras Mangueira CL, De Souza Barroso R, Rocha FA, Bacal NS, et al. The Use of Flow Cytometry in Myelodysplastic Syndromes: A Review. Front Oncol, 2017; 7: 270. doi: 10.3389/fonc.2017.00270.
- Choi JK, Xiao W, Chen X, Loghavi S, Elenitoba-Johnson KS, Naresh KN, et al. WHO 5th Edition Classification Project. Fifth Edition of the World Health Organization Classification of Tumors of the Hematopoietic and Lymphoid Tissues: Acute Lymphoblastic Leukemias, Mixed-Phenotype Acute Leukemias, Myeloid/Lymphoid Neoplasms with Eosinophilia, Dendritic/Histiocytic Neoplasms, and Genetic Tumor Syndromes. Mod Pathol, 2024; 37(5): 100466. doi: 10.1016/j.modpat.2024.100466.
- Fisher-Heath TC, Sharma A, Marshall MS, Brown T, Batra S. BCOR, BCORL1, and BCL6Mutations in Pediatric Leukemias. Cancers (Basel), 2025; 17(15): 2443. doi: 10.3390/cancers17152443.
- Yasuda T, Ueno T, Fukumura K, Yamato A, Ando M, Yamaguchi H, et al. Leukemic evolution of donor-derived cells harboring IDH2 and DNMT3A mutations after allogeneic stem cell transplantation. Leukemia, 2014; 28(2): 426-428. doi: 10.1038/leu.2013.278.
- Ihlow J, Penter L, Vuong LG, Bischoff P, Obermayer B, Trinks A, et al. Diagnosing recipient- vs. donor-derived posttransplant myelodysplastic neoplasm via targeted single-cell mutational profiling. Med, 2025; 6(4): 100548. doi: 10.1016/j.medj.2024.11.001.

