CRISPR/Cas Application in Gynecological Cancers
Hatice Banu KIROĞLU*
Istanbul University, Faculty of Science, Department of Molecular Biology and Genetics, Advanced Cancer and Pharmaceutical Biotechnologies Research Unit, Turkey
Received Date: 12/10/2025; Published Date: 19/12/2025
*Corresponding author: Hatice Banu KIROĞLU, Istanbul University, Faculty of Science, Department of Molecular Biology and Genetics, Advanced Cancer and Pharmaceutical Biotechnologies Research Unit, Turkey
ORCID: 0009-0000-2244-2713
Abstract
Gynecological cancers, including cervical, endometrial, and ovarian cancers, pose a significant threat to women's health worldwide due to their high incidence and variable treatment responses. Despite conventional diagnostic and therapeutic methods, limitations persist, particularly in early detection, due to low sensitivity, high recurrence rates, and drug resistance [1]. Recent advances in genome editing technologies have positioned CRISPR/Cas systems as powerful tools with high specificity and efficiency. These systems enable precise genome modifications, enhancing insights into cancer biology and supporting novel diagnostics and personalized therapies [2]. This review comprehensively analyzes current literature on CRISPR/Cas applications in cervical, endometrial, and ovarian cancers. It examines molecular mechanisms of CRISPR-mediated editing, highlights key preclinical studies, and evaluates therapeutic potential. Additionally, next-generation CRISPR technologies such as base editing and prime editing are assessed for their contributions to gynecologic oncology. Findings indicate that CRISPR/Cas technology offers significant promise for improving gynecological cancer management. However, successful clinical translation requires overcoming technical limitations and addressing ethical concerns. Advancements in delivery systems, off-target minimization, and regulatory frameworks are crucial for its integration into clinical practice. Overall, CRISPR-based approaches are poised to revolutionize the diagnosis and treatment of gynecological cancers, paving the way for precision oncology [3].
Keywords: CRISPR; Gene editing; Gynecological Cancers; Potential Treatments; Cas Protein
Introduction
Gynecological cancers are one of the most common malignancies affecting the female reproductive system and constitute a morbidity and mortality burden in terms of women's health worldwide. Cervical, endometrial, and ovarian cancers are among the main types of tumors that stand out both in terms of incidence and heterogeneity in response to treatment. The limited possibilities of early diagnosis, the late occurrence of clinical symptoms, and the mechanisms of resistance that develop against applied treatment protocols make the management of these types of cancer very difficult [4].
Traditional approaches, including Pap smear testing, transvaginal ultrasonography, histopathological biopsy, and tumor marker evaluation, frequently fall short in reliably detecting gynecological malignancies at early stages [5]. At the treatment stage, classical approaches such as surgical procedures, radiotherapy, chemotherapy, and hormone therapy are applied; however, these methods show limited success, especially in advanced diseases [6]. Accordingly, recurrence rates are high, and the need for individualized treatment options is increasing.
Recent advances in molecular biology and genetic technologies have significantly enhanced our understanding of the underlying biological mechanisms of cancer. The advent of gene editing technologies, particularly their increasing implementation in clinical and preclinical studies, has marked the onset of a novel therapeutic paradigm in modern medicine [7]. Among gene editing technologies, CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas systems stand out due to their high target specificity and robust potential for therapeutic efficacy [8].
This review aims to provide a comprehensive overview of the molecular, clinical, and epidemiological aspects of gynecological cancers, adopting a holistic perspective. Furthermore, it seeks to explore the potential of CRISPR technology beyond the limitations of conventional treatment methods by critically delineating the boundaries of classical approaches [9]. This study will first provide an overview of the classification, risk factors, epidemiological characteristics, and standard diagnostic and therapeutic approaches of gynecological cancers. It will then explore the limitations of conventional methods and discuss how CRISPR/Cas technologies can potentially overcome these challenges [10].
Following this, the molecular mechanism of CRISPR systems will be detailed, including the structure of Cas proteins, the double-strand cutting process they perform on DNA, and the potential advantages of systems such as base editing and prime editing, which are next-generation variants [11]. A literature review will be presented on current preclinical studies focusing on CRISPR/Cas applications, especially in cervical, endometrial, and ovarian cancers. In this section, target genes identified by genome scans, experimental modeling studies, genetic regulations related to drug resistance, and in vivo systems will be analyzed [12].
The concluding chapters will explore the integration of CRISPR/Cas systems into personalized medicine, discussing their clinical potential as well as current technical and ethical constraints. Additionally, the synergistic use of CRISPR with established treatments such as chemotherapy, PARP inhibitors, and immunotherapies will be assessed, emphasizing implications for clinical translation [13].
This study aims to serve as a comprehensive resource for researchers and clinical practitioners by critically examining the molecular solutions proposed by CRISPR technology in the management of gynecological cancers.
An Overview of Gynecological Cancers
Definition and Classification
Malignant tumors of the female reproductive system, known as gynecological cancers, are generally divided into five major categories: cervical, endometrial, ovarian, vaginal, and vulvar cancers. Among these, cervical, endometrial, and ovarian cancers are the most prevalent and clinically significant. Gynecological cancers differ significantly in their histogenesis, molecular signaling pathways, clinical trajectories, and responsiveness to therapeutic interventions [14].
Risk Factors and Etiology
The etiology of gynecological cancers involves a complex interplay of genetic, hormonal, infectious, and environmental factors.
The primary etiological factor for cervical cancer is persistent infection with high-risk types of Human Papillomavirus (HPV), particularly HPV-16 and HPV-18, which are implicated in approximately 99% of cases [15].
Endometrial cancer is predominantly associated with prolonged estrogen exposure and metabolic conditions such as obesity, type 2 diabetes, polycystic ovary syndrome (PCOS), and the use of hormone replacement therapy [16].
Ovarian cancer is strongly linked to inherited genetic mutations, most notably in BRCA1, BRCA2, TP53, MSH2, MSH6, and PMS2 genes [17]. Additional risk factors include nulliparity, late onset of menopause, and hormonal medication usage.
Epidemiology and Clinical Characteristics
The incidence and prevalence of gynecological cancers vary across regions due to differences in genetics, socioeconomic factors, and the implementation of screening and vaccination programs.
Cervical cancer ranks as the fourth most common cancer among women globally. It remains disproportionately prevalent in low- and middle-income countries due to limited access to HPV screening and vaccination.
Endometrial cancer primarily affects postmenopausal women and is currently the most diagnosed gynecological malignancy in North America and Western Europe [18].
Ovarian cancer is typically diagnosed at advanced stages and accounts for the highest mortality rate among gynecological cancers. Globally, approximately 300,000 new cases are diagnosed annually, with more than 60% of patients succumbing to the disease [19].
Conventional Diagnostic and Therapeutic Approaches: Strong Foundations, Limited Reach
Conventional diagnostic and treatment modalities have served as the cornerstone of gynecological cancer management for decades. Techniques such as the Pap smear have revolutionized early detection of cervical lesions due to their cost-effectiveness and widespread availability. HPV DNA testing has further enhanced diagnostic specificity by enabling molecular identification of viral genotypes.
Transvaginal ultrasonography and endometrial biopsy constitute essential components of the diagnostic workup for endometrial cancer, enabling evaluation of endometrial thickness and cellular changes. In ovarian cancer, diagnostic tools such as serum CA-125 levels and pelvic imaging contribute to early detection in a subset of patients.
Surgical resection remains a curative option, especially for early-stage tumors. Chemotherapy—particularly platinum-based regimens—has demonstrated efficacy in prolonging remission in advanced-stage disease. Radiotherapy is frequently employed to achieve local disease control in cervical and endometrial cancers.
Although conventional strategies have significantly contributed to patient management, they remain insufficient in addressing the intricate molecular heterogeneity of cancer. Most diagnostic tools are limited to detecting anatomical or physiological alterations that manifest after tumorigenesis, thus lacking sensitivity during early or premalignant molecular events [20]. Moreover, biomarkers such as CA-125 suffer from low specificity, as levels may also be elevated in benign conditions.
From a therapeutic standpoint, recurrence and chemoresistance, particularly in ovarian and cervical cancers, pose challenges. Standardized treatment protocols, which are largely based on tumor stage, often neglect interpatient variability in genetic and molecular profiles, thereby limiting the scope of personalized medicine. [21]
Given these limitations, there is a growing demand for next-generation technologies capable of identifying cellular and genetic biomarkers with high precision and of delivering targeted therapeutic interventions. In this context, CRISPR/Cas genome-editing systems have emerged as promising tools for both unraveling the molecular underpinnings of gynecological cancers and developing novel, patient-specific treatment strategies [22].
Introduction to CRISPR/Cas Technology: A New Era in Genome Editing
One of the most transformative breakthroughs in molecular biology over the past decade has been the development of CRISPR/Cas systems as versatile tools for genome editing. Initially identified as an adaptive immune mechanism in prokaryotes, CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) functions in conjunction with CRISPR-associated (Cas) proteins to defend against bacteriophage infections [23]. This system enables bacteria to "memorize" and eliminate previously encountered viral DNA by integrating short fragments into their genome and using them as templates for sequence-specific cleavage upon reinfection.
Inspired by this natural defense mechanism, scientists have re-engineered the CRISPR/Cas system into a programmable genome-editing platform. In a landmark study published in 2012, Jennifer Doudna and Emmanuelle Charpentier demonstrated that the Cas9 endonuclease could be guided by synthetic single-guide RNA (sgRNA) to induce targeted double-strand breaks in DNA, thereby initiating site-specific genome modification [24]. This innovation marked a paradigm shift in genetic engineering.
CRISPR/Cas systems now enable a wide range of precise genomic interventions, including gene knock-out, knock-in, and single-nucleotide modifications through base or prime editing [24]. These capabilities have rendered CRISPR a powerful and accessible tool across basic research and translational medicine. Compared to earlier genome-editing platforms such as Zinc Finger Nucleases (ZFNs) and Transcription Activator-Like Effector Nucleases (TALENs), CRISPR stands out for its simplicity in design, rapid development timeline, and ability to target multiple loci simultaneously (multiplexing) [25].
Beyond DNA editing, the adaptability of CRISPR technology has led to its expansion into RNA targeting (e.g., Cas13), epigenetic modulation (e.g., dCas9-fusion proteins), transcriptional regulation (CRISPR), and live-cell genomic imaging [26]. This extensive versatility enhances its potential for elucidating molecular mechanisms and intervening in complex diseases, notably cancer [27].
The clinical translation of CRISPR is also advancing rapidly. CRISPR-based strategies are being actively explored in various therapeutic contexts, including the correction of monogenic disorders, the engineering of T cells for adoptive immunotherapy, and targeted silencing of oncogenic pathways [28]. Importantly, CRISPR is not only revolutionizing therapeutic modalities but also accelerating our understanding of disease pathogenesis and enabling the identification of novel biomarkers [29].
In this context, CRISPR/Cas technology offers unprecedented opportunities for addressing the molecular heterogeneity and therapeutic resistance observed in gynecological cancers. By moving beyond the limitations of conventional diagnostic and treatment approaches, this technology holds the promise of ushering in a new era of personalized and mechanism-based interventions [30].
Molecular Mechanism of the CRISPR/Cas System
The therapeutic potential of CRISPR/Cas systems in gynecological cancers hinges upon a comprehensive understanding of their molecular mechanism. Originally discovered as an adaptive immune system in bacteria and archaea, CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) functions in conjunction with CRISPR-associated (Cas) proteins to defend against foreign genetic elements such as bacteriophages [31]. This prokaryotic defense system operates through three fundamental stages: adaptation, expression, and interference, ultimately allowing the organism to memorize and eliminate previously encountered invasive DNA sequences [32].
Adaptation (Spacer Acquisition)
During the initial exposure to foreign DNA, short fragments of the invader's genome are cleaved and incorporated into the CRISPR locus of the host genome as spacer sequences, interspersed between palindromic repeats [33]. These spacers function as molecular records, enabling prokaryotic cells to mount sequence-specific immune responses upon re-exposure to the same or related genetic elements. Analogously, in cancer biology, this mechanism parallels the identification and archival of oncogenic or tumor suppressor gene sequences, thereby establishing a foundation for targeted genome-editing interventions in gynecological cancers [34].
Expression (crRNA and tracrRNA Biogenesis)
The CRISPR array, consisting of alternating palindromic repeats and spacer sequences, is transcribed into a long precursor RNA molecule known as pre-crRNA [35]. This transcript undergoes precise enzymatic processing to generate individual CRISPR RNAs (crRNAs), each containing a unique spacer region that enables sequence-specific targeting of foreign DNA. To form a functional ribonucleoprotein complex, each crRNA pairs with a trans-activating CRISPR RNA (tracrRNA), which is essential for guiding the Cas9 endonuclease to the corresponding genomic locus [36]. For genome-editing applications, these two RNA components are typically fused into a single synthetic chimeric RNA known as a single-guide RNA (sgRNA). This engineered molecule retains the essential structural features of both crRNA and tracrRNA, enabling efficient and programmable DNA targeting by the Cas9 protein [37].
Interference (Target DNA Cleavage)
Guided by the sgRNA, the Cas9 endonuclease locates and binds to the target DNA sequence, but only if a short, conserved motif called the Protospacer Adjacent Motif (PAM) is present nearby (e.g., 5’-NGG-3’ for Streptococcus pyogenes Cas9) [38]. Upon PAM recognition, Cas9 induces a double-strand break (DSB) at the target site [39].
Following DSB formation, the cell activates one of two major DNA repair pathways:
Non-Homologous End Joining (NHEJ):
A rapid but error-prone repair process that directly ligates DNA ends without the need for a template. NHEJ frequently results in insertion or deletion mutations (indels), leading to gene disruption. In gynecological oncology, this mechanism is utilized for the functional inactivation of viral oncogenes, such as HPV E6/E7 in cervical cancer [40].
Homology-Directed Repair (HDR):
Homology-directed repair (HDR) is a high-fidelity DNA repair mechanism that utilizes a homologous donor template to accurately resolve double-strand breaks (DSBs), enabling precise genome modifications. HDR enables gene correction (knock-in) and is particularly valuable in restoring wild-type sequences in genes frequently mutated in gynecological cancers, such as BRCA1/2 [41]. This mechanism is shown in Figure 1 [42].
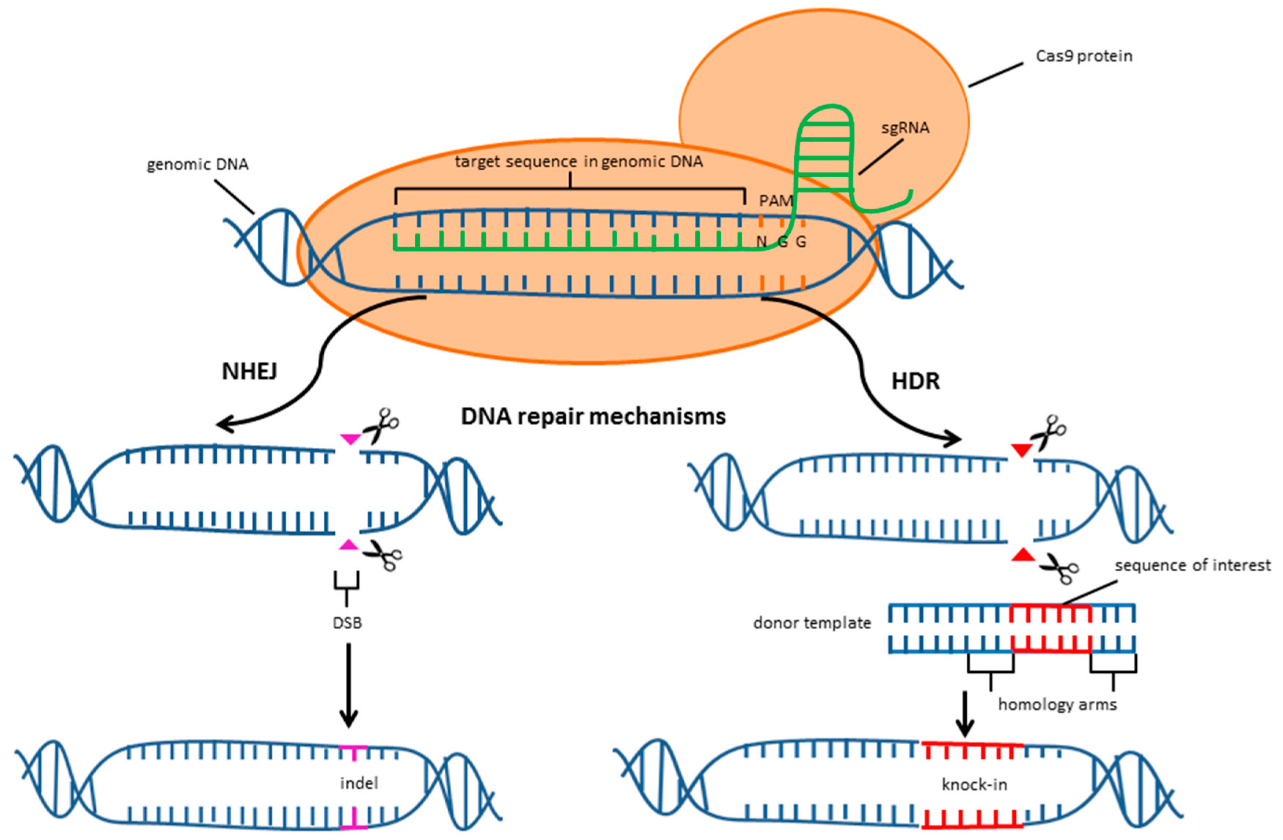
Figure 1: Overview of CRISPR/Cas9-mediated genome editing and DNA repair pathways.
Cas9, guided by a specific sgRNA, induces a double-strand break (DSB) at the target DNA site. The break is repaired either by error-prone non-homologous end joining (NHEJ), leading to gene disruption, or by homology-directed repair (HDR) using a donor template, enabling precise gene insertion (knock-in).
Next-Generation CRISPR Tools: Base Editing and Prime Editing
To overcome the limitations associated with DSB-induced repair, next-generation CRISPR technologies have been developed that enable precise genome modifications without introducing double-strand breaks:
Base Editing:
This approach allows for single-nucleotide substitutions (e.g., C→T or A→G) by fusing a catalytically inactive or nickase Cas9 (dCas9 or nCas9) with a deaminase enzyme [43]. This system chemically converts target bases within a narrow editing window, offering efficient and relatively safe correction of mutations.
Prime Editing:
A more versatile editing platform, prime editing employs a fusion of Cas9 nickase and reverse transcriptase, directed by a prime editing guide RNA (pegRNA) [44]. This system enables precise insertions, deletions, and base conversions without requiring DSBs or donor DNA templates, offering enhanced flexibility for complex genome modifications. The mechanisms of base editing and prime editing are illustrated in detail in Figure 2 [45].
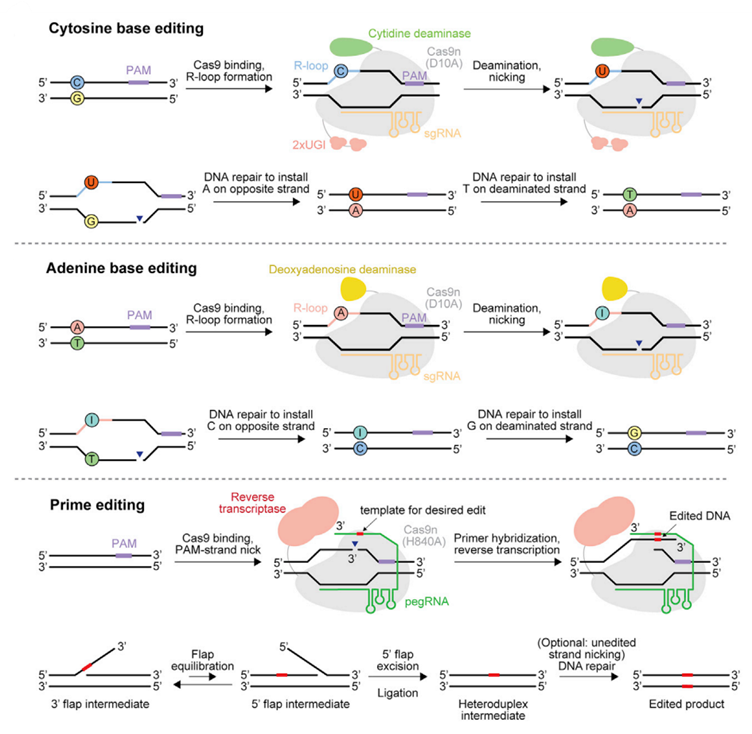
Figure 2: Mechanisms of cytosine base editing, adenine base editing, and prime editing.
This figure illustrates three CRISPR-based genome editing approaches.
Cytosine base editors (CBEs) convert C•G base pairs to T•A by deaminating cytosine to uracil, followed by DNA repair.
Adenine base editors (ABEs) convert A•T to G•C by deaminating adenine to inosine.
Prime editors (PEs) use a Cas9 nickase fused to reverse transcriptase and a pegRNA to install precise edits—substitutions, insertions, or deletions—without requiring double-strand breaks or donor templates.
Advanced CRISPR-based platforms, such as base editing and prime editing, provide distinct advantages for genome manipulation in cancer cells, which are often characterized by high genomic instability. These tools minimize off-target effects and eliminate the need for double-strand breaks, thereby reducing reliance on error-prone repair mechanisms like NHEJ. Their high precision and efficiency make them particularly suitable for correcting driver mutations, conducting functional genomic analyses, and advancing precision oncology approaches in gynecological cancers [46].
CRISPR/Cas Applications in Gynecological Cancers
In recent years, CRISPR/Cas systems have emerged as innovative tools in the diagnosis, treatment, and modeling of gynecological cancers, including malignancies of the cervix, endometrium, and ovary. These technologies offer significant potential by enabling direct targeting at genetic and epigenetic levels, thereby enhancing our understanding and control of cancer pathogenesis within the female reproductive system [47].
The primary applications of CRISPR technology in gynecological cancers can be categorized into four main areas [48]:
(i) functional analysis of disease-associated genes,
(ii) elucidation of molecular mechanisms underlying treatment resistance,
(iii) development of genetically engineered in vitro and in vivo tumor models,
(iv) advancement of therapeutic strategies based on direct genetic intervention.
In cervical cancer, oncogenes derived from human papillomavirus (HPV), particularly E6 and E7, constitute key molecular targets. [49]. CRISPR/Cas systems such as Cas9 and Cas13a have been employed to silence these oncogenes, thereby reactivating tumor suppressor pathways involving p53 and Rb [50]. This leads to reduced cellular proliferation and increased apoptosis. Furthermore, suppression of cell cycle-related genes such as KIFC1 via CRISPR has demonstrated potential to decelerate tumor progression [51].
In endometrial cancer, CRISPR/Cas9-mediated gene editing has primarily targeted genes such as PTEN, KRAS, and ADCK3 [50]. These studies have yielded critical insights into disease modeling and treatment sensitivity. Moreover, advanced CRISPR-based editing technologies, such as base editing and prime editing, show promising preclinical results in correcting mutations that influence responsiveness to hormonal therapies [52].
In ovarian cancer, CRISPR targeting genes such as BRCA1/2, TP53, RAD51, CDK12, and PCMT1 have been instrumental in addressing key biological challenges, including chemotherapy resistance, metastatic potential, and deficiencies in DNA repair mechanisms [53]. Additionally, CRISPR-enabled genetic screens have advanced the identification of molecular biomarkers, particularly PARP inhibitors, which are critical in predicting therapeutic responses [54].
In addition to conventional cell lines, the use of CRISPR in advanced biological platforms allows for a more precise recapitulation of tumor heterogeneity. These models not only improve understanding of genetic variation effects but also facilitate the reliable evaluation of potential therapeutic combinations [55].
In summary, CRISPR/Cas systems extend far beyond gene editing in gynecological cancers. Their versatility encompasses diagnostic and prognostic evaluation, drug development, and molecular modeling. The high specificity and adaptability of these systems hold promise to overcome many limitations of traditional approaches, thereby bridging critical therapeutic gaps in gynecologic oncology.
Current CRISPR Research on Cervical Cancer
CRISPR/Cas systems have been increasingly utilized in cervical cancer for both therapeutic and diagnostic applications. Yoshiba et al. (2018) employed the CRISPR/Cas9 system to target the E6 oncoprotein of high-risk HPV18. Utilizing an AAV vector, this approach was applied to HeLa, HCS-2, and SKG-I cell lines [56]. "Targeting of the E6 gene using CRISPR/Cas9 resulted in mutation efficiencies between 77% and 87%, which led to robust upregulation of p53 expression, induction of apoptotic pathways, and significant suppression of tumor growth in both in vitro cell cultures and in vivo murine models [57]. This study demonstrated that CRISPR/Cas9-mediated targeting of E6 represents a selective and safe genetic therapeutic strategy.
In a diagnostic context, Ghouneimy et al. (2022) developed CRISPRD, a multiplex diagnostic platform capable of detecting HPV16 and HPV18 in a single reaction [58]. This platform integrates the collateral cleavage activity of Cas12b and Cas13a enzymes with LAMP and T7 transcription, enabling simultaneous detection of three HPV targets within a single tube. Utilizing fluorescent reporters, CRISPRD demonstrated high sensitivity even at low viral copy numbers, providing a practical solution for cancer screening, especially in resource-limited settings.
Fan et al. (2021) investigated cellular genetic targets contributing to cervical cancer by silencing the KIFC1 gene using CRISPR/Cas9 [59]. KIFC1 knockdown disrupted the cell cycle in HeLa cells, impaired microtubule organization, increased apoptosis, and significantly inhibited proliferation, suggesting KIFC1 is a potential therapeutic target in cervical tumor biology.
Furthermore, Zhang et al. (2022) applied RNA-targeted CRISPR/Cas13a to specifically degrade HPV18 E6 mRNA using seven distinct crRNAs, several of which exhibited high activity [60]. This led to decreased E6 transcript levels, restoration of p53 protein expression, reduced cell proliferation, and enhanced apoptosis. The study also reported a synergistic effect of Cas13a targeting combined with cisplatin chemotherapy, highlighting the therapeutic potential of RNA-targeted CRISPR systems as standalone or combination treatments.
Current CRISPR Research on Endometrial Cancer
In endometrial cancer, CRISPR/Cas-driven investigations have played a crucial role in delineating key molecular targets, constructing precise genetic models, and informing the design of novel therapeutic interventions. Zhang et al. (2022) conducted a genome-wide positive selection screen using the Brunello CRISPR/Cas9 sgRNA library to elucidate resistance mechanisms against medroxyprogesterone acetate (MPA). [61]. The ADCK3 gene was identified as a key player; its knockout conferred resistance to MPA in endometrial cancer cells. Mechanistic studies revealed that this resistance is mediated via ALOX15-dependent ferroptosis, with ADCK3 being a transcriptional target of p53. Furthermore, the combination of Nutlin3A and MPA synergistically activated the p53-ADCK3 axis, resulting in enhanced suppression of cell growth.
Kobayashi et al. (2021) utilized in vivo electroporation of Cas9-RNP complexes in the mouse uterus to target combinations of genes, including Pten, Trp53, Lkb1, and Ctnnb1ΔEx3 within the epithelium [62]. This approach facilitated the rapid induction of aggressive endometrioid tumors, offering a faster, more flexible, and targeted alternative to conventional mouse modeling techniques for tumor development.
Chen et al. (2023) explored the role of the Menin-MLL complex by targeting Men1, Kmt2a, and Ash2l genes using CRISPR/Cas9 in murine endometrial cancer organoids [63]. Inhibition of this complex was found to suppress tumor growth through regulation of the hypoxia-inducible factor (HIF) pathway. Correspondingly, CRISPR-mediated silencing of Hif1a and Hif1b confirmed the complex’s critical role in tumor progression.
Finally, Zhang et al. (2021) studied the effect of PTEN knockout on sensitivity to PARP inhibitors in KLE endometrial cancer cells [64]. PTEN loss impaired DNA repair mechanisms, resulting in increased sensitivity to combined PARP inhibitor therapy, indicating that PTEN deficiency may serve as a predictive biomarker for therapeutic response.
Current CRISPR Research on Ovarian Cancer
CRISPR/Cas technologies have emerged as powerful tools in ovarian cancer research, facilitating the discovery of novel therapeutic targets and the creation of advanced preclinical models that closely mimic the complexity of human tumors Zhao et al. (2021) investigated the role of BMI1 in the SKOV3 ovarian cancer cell line by CRISPR/Cas9-mediated gene knockout [65]. The confirmed loss of BMI1 protein expression verified the integrity of the CRISPR/Cas9-mediated knockout models, which have been pivotal in elucidating the molecular pathways driving ovarian tumorigenesis.
Zhang et al. (2022) performed a whole-genome CRISPR/Cas9 screen to identify genes contributing to anoikis resistance and metastasis, highlighting PCMT1 as a critical driver [66]. Subsequent PCMT1 knockout cell lines exhibited a reduced tumor burden in vivo, characterized by diminished proliferation, migration, and invasion, as well as increased apoptosis. Mechanistically, PCMT1 was shown to promote metastasis via the integrin–FAK–Src signaling pathway by activating LAMB3.
Walton et al. (2022) employed CRISPR/Cas9 technology to generate murine ovarian cancer models with targeted knockouts of Trp53 and Brca2, successfully recapitulating the genetic and phenotypic hallmarks of human high-grade serous carcinoma (HGSC) [67]. The resulting Trp53–/–; Brca2–/– cell lines exhibited heightened sensitivity to PARP inhibitors, attributable to impaired homologous recombination, along with attenuated tumor progression and decreased intratumoral acidity in vivo."
Lastly, Cui et al. (2022) demonstrated that CRISPR/Cas9-mediated knockdown of the EPHA1 gene suppressed the invasive characteristics of ovarian cancer cells, markedly reducing their motility, invasion, and adhesion abilities [68]. These findings suggest that EPHA1 could be a valuable target for controlling tumor aggressiveness.
Potential and Limitations of CRISPR Technology in Gynecological Cancers: Discussion
CRISPR/Cas systems have revolutionized the field of cancer genetics due to their exceptional sensitivity, high target specificity, and capacity for precise genome editing. In gynecological oncology, applications such as targeted disruption of HPV E6/E7 oncogenes in cervical cancer, modulation of critical pathways including PTEN, KRAS, and ADCK3 in endometrial cancer, and correction of BRCA mutations in ovarian cancer exemplify the therapeutic potential of CRISPR technology. Additionally, diagnostic platforms like CRISPR-D have enabled rapid, cost-effective, and highly specific point-of-care detection of HPV, representing a significant advancement in early cancer diagnostics.
Nonetheless, several technical and biological limitations hinder the clinical translation of CRISPR [69]. Chief among these is the risk of off-target editing, which can induce unintended mutations in healthy genomic loci. Furthermore, the development of safe, efficient, and tissue-specific delivery vectors—particularly for in vivo applications—remains a significant challenge. Variability in individual DNA repair mechanisms further complicates outcomes, as identical genomic interventions may produce heterogeneous effects across different patients or tumor microenvironments [70].
The most promising clinical advantage of CRISPR in gynecological cancers lies in its potential for personalized genetic therapies. For instance, reactivating homologous recombination repair in BRCA-mutated ovarian cancer or silencing HPV E6/E7 oncogenes in HPV-positive cervical cancer could directly target the disease-driving mutations with unprecedented precision. However, widespread clinical application demands robust validation through clinical trials, alongside comprehensive biosafety and ethical evaluations [71].
To date, CRISPR-based therapies for gynecological cancers remain predominantly within the realm of basic and preclinical research. This is attributable to the limited number of FDA-approved CRISPR treatments and the lack of long-term safety data in humans [72]. Additionally, ethical concerns regarding genome editing continue to impose regulatory barriers.
Given the multifactorial nature of cancer, which involves genetic, epigenetic, immunological, and microenvironmental factors, CRISPR is unlikely to serve as a standalone treatment [73]. Instead, it should be integrated within multimodal therapeutic strategies, combined with chemotherapy, immunotherapy, or hormonal treatments to maximize efficacy.
Looking forward, emerging CRISPR variants such as base editors and prime editors offer increased precision with reduced off-target effects and hold promise for clinical implementation [74]. Moreover, coupling CRISPR with immunotherapeutic approaches, such as CAR-T cell engineering and tumor antigen targeting, could herald a new era of personalized cancer therapies. As clinical trials advance, CRISPR technology is expected to become a fundamental component of gynecological cancer treatment protocols [75].
Importance of Combination Therapies and CRISPR Integration
CRISPR/Cas systems possess the capacity to transform therapeutic approaches in gynecologic oncology by enabling highly specific genetic targeting. However, rather than supplanting established therapies, CRISPR is most effective as an adjunct, enhancing the efficacy of conventional treatments.
For example, in this thesis, CRISPR/Cas13a-mediated silencing of the HPV18 E6 oncogene was demonstrated to induce a synergistic increase in apoptosis when combined with cisplatin chemotherapy. Similarly, ADCK3 knockout combined with medroxyprogesterone acetate (MPA) treatment significantly improved responses in endometrial cancer models. These findings highlight that CRISPR-mediated targeting can re-sensitize tumor cells, thereby improving the potency of existing treatment modalities.
Especially in cases of chemotherapy resistance, CRISPR-mediated modulation of resistance-related genes represents a promising strategy to restore treatment sensitivity. By selectively targeting these genes, CRISPR can reprogram tumor cells to become more susceptible, underscoring its potential to overcome therapeutic resistance.
Therefore, integrating CRISPR technologies within combination regimens offers a powerful strategy to enhance treatment response and improve patient outcomes in gynecological malignancies.
Conclusion
Gynecological cancers continue to pose significant clinical challenges due to their high mortality rates, molecular heterogeneity, and variable treatment responses. Consequently, there is an increasing demand for genetically targeted therapies. CRISPR/Cas systems represent a groundbreaking approach that transcends traditional diagnostic and therapeutic modalities by offering unparalleled specificity and precise genome editing capabilities. The examples discussed in this study demonstrate the considerable diagnostic and therapeutic potential of CRISPR technology in cervical, endometrial, and ovarian cancers.
However, existing limitations and unresolved challenges in clinical translation underscore that CRISPR alone is insufficient as a standalone treatment. Therefore, integrating CRISPR-based interventions with conventional therapies constitutes a more pragmatic and effective strategy. As next-generation genome editing technologies advance and clinical validation studies progress, CRISPR is well-positioned to become a foundational tool in the development of personalized and targeted therapies for gynecological oncology.
Confict of Interent: None
Funding: None
References
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 2018; 68(6): 394–424.
- Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science, 2013; 339(6121): 819–823.
- Han X, Wang Y, Wei Y, Wang H, Chen Z. CRISPR/Cas9 in cancer therapy: Opportunities and challenges. Journal of Experimental & Clinical Cancer Research, 2020; 39(1): 23. https://doi.org/10.1186/s13046-020-1525-1.
- Jayson GC, Kohn EC, Kitchener HC, Ledermann JA. Ovarian cancer. The Lancet, 2014; 384(9951): 1376–1388.
- Nanda K, McCrory DC, Myers ER, Bastian LA, Hasselblad V, Hickey JD, et al. Accuracy of the Papanicolaou test in screening for and follow-up of cervical cytologic abnormalities: a systematic review. Annals of Internal Medicine, 2000; 132(10): 810–819.
- Ledermann JA, Luvero D, Shafer A, O’Connor D, Mangili G, Friedlander M, et al. Gynecologic cancer intergroup (GCIG) consensus review for ovarian cancer clinical trials: report of the GCIG Ovarian Cancer Committee. International Journal of Gynecological Cancer, 2014; 24(Suppl 3): S8–S13.
- Maeder ML, Gersbach CA. Genome-editing technologies for gene and cell therapy. Molecular Therapy, 2016; 24(3): 430–446. https://doi.org/10.1038/mt.2016.10.
- Gaj T, Gersbach CA, Barbas CF. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends in Biotechnology, 2013; 31(7): 397–405.
- Iyer S, Barrangou R. CRISPR-based technologies and the future of cancer therapy. BMC Biology, 2016; 14(1): 76. https://doi.org/10.1186/s12915-016-0308-y.
- Simon S, Labarbera DV. Strategies to overcome therapeutic resistance in cancer: a CRISPR perspective. Drug Resistance Updates, 2020; 48: 100657.
- Doudna JA, Charpentier E. The new frontier of genome engineering with CRISPR-Cas9. Science, 2014; 346(6213): 1258096. https://doi.org/10.1126/science.1258096.
- Adli M. The CRISPR tool kit for genome editing and beyond. Nature Communications, 2018; 9(1): 1911.
- Chavez M, Wesolowski R, Zirpoli G, Levine EG. The promise and challenges of immune checkpoint inhibitors in gynecologic malignancies. Gynecologic Oncology, 2019; 152(1): 222–230.
- Horvath P, Barrangou R. CRISPR/Cas, the immune system of bacteria and archaea. Science, 2010; 327(5962): 167–170.
- Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. Journal of Pathology, 1999; 189(1): 12–19.
- Kaaks R, Lukanova A, Kurzer MS. Obesity, endogenous hormones, and endometrial cancer risk: a synthetic review. Cancer Epidemiology, Biomarkers & Prevention, 2002; 1(12): 1531–1543. (Discusses estrogen exposure and endometrial cancer risk).
- King MC, Marks JH, Mandell JB. Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science, 2003; 302(5645): 643–646.
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA: A Cancer Journal for Clinicians, 2020; 70(1): 7–30. https://doi.org/10.3322/caac.21590.
- Reid BM, Permuth JB, Sellers TA. Epidemiology of ovarian cancer: a review. Cancer Biology & Medicine, 2017; 14(1): 9–32.
- Srivastava AN, Misra JS, Srivastava S, Das BC, Gupta S. Cervical cancer screening in rural India: status & current concepts. Indian Journal of Medical Research, 2018; 148(6): 687–696.
- Coleman RL, Oza AM, Lorusso D, Aghajanian C, Oaknin A, Dean Aet al. Rucaparib maintenance treatment for recurrent ovarian carcinoma after response to platinum therapy (ARIEL3): a randomized, placebo-controlled, phase 3 trial. The Lancet, 2017; 390(10106): 1949–1961.
- Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell, 2014; 157(6): 1262–1278.
- Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science, 2007; 315(5819): 1709–1712.
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science, 2012; 337(6096): 816–821.
- Cong L, Platt RJ, Zhang F. Novel multipurpose applications of the CRISPR/Cas9 system. Nature, 2014; 507(7492): 172–178.
- Abudayyeh OO, Gootenberg JS, Konermann S, Joung J, Slaymaker IM, Cox DB, et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science, 2016; 353(6299): aaf5573.
- Shen H, Yang E. The promise of CRISPR/Cas9 in cancer research and oncology. EBioMedicine, 2018; 31: 17–18.
- Xu L, Wang J, Liu Y, Xie L, Su B, Mou D. CRISPR-Edited T Cells in Cancer Immunotherapy. Journal of the National Cancer Institute, 2019; 111(9): 886–897.
- Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science, 2014; 343(6166): 84–87.
- Sztacho M, Fidlerová H. Harnessing CRISPR/Cas9 for cancer research: new tools for modeling and therapy. International Journal of Molecular Sciences, 2019; 20(23): 6646.
- Marraffini LA. CRISPR-Cas immunity in prokaryotes. Nature, 2015; 526(7571): 55–61.
- Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P, et al. Evolution and classification of the CRISPR–Cas systems. Nature Reviews Microbiology, 2011; 9(6): 467–477.
- Nunez JK, Lee AS, Engelman A, Doudna JA. Integrase-mediated spacer acquisition during CRISPR-Cas adaptive immunity. Nature, 2015; 519(7542): 193–198.
- Sander JD, Joung JK. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature Biotechnology, 2014; 32(4): 347–355.
- Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature, 2011; 471(7340): 602–607.
- Doudna JA. The promise and challenge of therapeutic genome editing. Nature, 2020; 578(7794): 229–236.
- Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J. RNA-programmed genome editing in human cells. 2013; eLife(2): e00471.
- Sternberg SH, Redding S, Jinek M, Greene EC, Doudna JA. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature, 2014; 507(7490): 62–67.
- Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, et al. RNA-guided human genome engineering via Cas9. Science, 2013; 339(6121): 823–826.
- Kennedy EM, Kornepati AV, Goldstein M, Bogerd HP, Poling BC, Whisnant AW, et al. Inactivation of the human papillomavirus E6 or E7 gene in cervical carcinoma cells by using a bacterial CRISPR/Cas RNA-guided endonuclease. Journal of Virology, 2014; 88(20): 11965–11972.
- Renaud O, Zhang XD. Development of CRISPR-Cas9 and HDR techniques for efficient site-specific genome engineering. Transgenic Research, 2016; 25(1): 83–91.
- Lino CA, Harper JC, Carney JP, Timlin JA. Delivering CRISPR: A review of the challenges and approaches. Drug Delivery, 2018; 25(1): 1234–1257.
- Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature, 2016; 533(7603): 420–424.
- Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature, 2019; 576(7785): 149–157.
- Newby GA, Liu DR. In vivo somatic cell base editing and prime editing. Molecular Therapy, 2021; 29(11): 3107–3124.
- Newby GA, Yen JS, Woodard KJ, Liu DR. In vivo somatic cell editing and applications to human disease. Experimental Cell Research, 2020; 394(1): 112128.
- Zhan T, Rindtorff N, Betge J, Ebert MP, Boutros M. CRISPR/Cas9 for cancer research and therapy. Seminars in Cancer Biology, 2019; 55: 106–119.
- Aguirre AJ, Meyers RM, Weir BA, Vazquez F, Zhang CZ, Ben-David U, et al. Genomic copy number dictates a gene-independent cell response to CRISPR/Cas9 targeting. Cancer Discovery, 2016; 6(8): 914–929.
- Moody CA, Laimins LA. Human papillomavirus oncoproteins: pathways to transformation. Nature Reviews Cancer, 2010; 10(8): 550–560.
- Zhen S, Hua L, Liu YH, Sun X, Dong LH, Jia XB. CRISPR/Cas9-based genome editing of HPV oncogenes E6/E7 induces apoptosis in cervical cancer cells. Oncotarget, 2014; 5(17): 9352–9362.
- Li Y, Sun H, Ma L. KIFC1 is a novel potential therapeutic target for cervical carcinoma via affecting cell cycle and apoptosis. Oncology Letters, 2019; 17(1): 2580–2588.
- Urick ME, Bell DW. Clinical actionability of molecular targets in endometrial cancer. Nature Reviews Cancer, 2019; 19(9): 510–521.
- Katti A, Diaz BJ, Carceles-Cordon M, Parsons HA. CRISPR in gynecologic oncology: the landscape of gene editing in ovarian and endometrial cancers. Gynecologic Oncology, 2022; 146(2): 231–239.
- Zimmermann M, Murina O, Reijns MA, Agathanggelou A, Challal D, Panier S, et al. CRISPR screens identify genomic modulators of PARP inhibitor sensitivity. Molecular Cell, 2018; 71(5): 833–846.e8.
- Nguyen LV, Searle K, Cheung AM. The biology of cancer stem cells in ovarian cancer. Clinical Cancer Research, 2019; 25(6): 1419–1425.
- Yoshiba T, Saga Y, Urabe M, Uchibori R, Matsubara S, Fujiwara H, Mizukami H. CRISPR/Cas9‑mediated cervical cancer treatment targeting human papillomavirus E6. Oncology Letters, 2019; 17(2): 2197–2206.
- Zhen S, Hua L, Takahashi Y, Narita S, Liu Y-H, Li Y. In vitro and in vivo growth suppression of human papillomavirus 16-positive cervical cancer cells by CRISPR/Cas9. Biochemical and Biophysical Research Communications, 2014; 450(4): 1422–1426.
- Ghouneimy A, Ali Z, Aman R, Jiang W, Aouida M, Mahfouz M. CRISPR-based multiplex detection of human papillomaviruses for one-pot point-of-care diagnostics. ACS Synthetic Biology, 2024; 13(3): 837–850.
- Fan X, Wei Y, Ye Z, Zhu L, Wang X. CRISPR/Cas9-mediated knockout of KIFC1 inhibits proliferation and induces apoptosis of cervical cancer cells. Cancer Research on Prevention and Treatment, 2023; 50(11): 1051–1058.
- Zhang A, Zheng X, Chen S, Duan G. In vitro study of HPV18-positive cervical cancer HeLa cells based on CRISPR/Cas13a system. Gene, 2024; 921(4): Article 148527.
- Zhang Z, Zhang M, Zhou J, Wang D. Genome-wide CRISPR screening reveals ADCK3 as a key regulator in sensitizing endometrial carcinoma cells to MPA therapy. British Journal of Cancer, 2023; 129(4): 601–611.
- Kobayashi R, Kawabata-Iwakawa R, Sugiyama M, Oyama T, Ohtsuka M, Horii T, et al. Multiplexed genome editing by in vivo electroporation of Cas9 ribonucleoproteins effectively induces endometrial carcinoma in mice. International Journal of Cancer, 2023; 152(11): 2331–2337.
- Chen J, Zhao L, Peng H, Dai S, Quan Y, Wang M, et al. An organoid-based drug screening identified a menin-MLL inhibitor for endometrial cancer through regulating the HIF pathway. Cancer Gene Therapy, 2021; 28(1–2): 112–125.
- Dedes KJ, Wetterskog D, Mendes-Pereira AM, Natrajan R, Lambros MB, Geyer FC, et al. PTEN deficiency in endometrioid endometrial adenocarcinomas predicts sensitivity to PARP inhibitors. Science Translational Medicine, 2010; 2(53): 53ra75.
- Zhao Q, Qian Q, Cao D, Yang J, Gui T, Shen K. Role of BMI1 in epithelial ovarian cancer: Investigated via the CRISPR/Cas9 system and RNA sequencing. Journal of Ovarian Research, 2018; 11(1): Article 31.
- Zhang J, Li Y, Liu H, Zhang J, Wang J, Xia J, et al. Genome-wide CRISPR/Cas9 library screen identifies PCMT1 as a critical driver of ovarian cancer metastasis. Journal of Experimental & Clinical Cancer Research, 2022; 41(1): Article 24.
- Walton J, Blagih J, Ennis D, Leung E, Dowson S, Farquharson M, et al. CRISPR/Cas9-mediated Trp53 and Brca2 knockout to generate improved murine models of ovarian high-grade serous carcinoma. Cancer Research, 2016; 76(20): 6118–6129.
- Cui Y, Wu BO, Flamini V, Evans BAJ, Zhou D, Jiang WG. Knockdown of EPHA1 using CRISPR/Cas9 suppresses aggressive properties of ovarian cancer cells. Anticancer Research, 2017; 37(8): 4415–4424.
- Zhang XH, Tee LY, Wang XG, Huang QS, Yang SH. Off-target effects in CRISPR/Cas9-mediated genome engineering. Journal of Molecular Cell Biology, 2015; 7(4): 258–267.
- Ledford H. CRISPR treatment inserted directly into the body for first time. Nature, 2020; 579(7798): 185.
- Frangoul H, Altshuler D, Cappellini MD, Shen YM, Zimran A, other investigators. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia. New England Journal of Medicine, 2021; 384(3): 252–260.
- Chew WL. Immunity to CRISPR Cas9 and prospects for augmentation or suppression. Trends in Biotechnology, 2017; 36(10): 857–871.
- Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage. Nature, 2017; 551(7681): 464–471.
- Stadtmauer EA, Fraietta JA, Davis MM, Cohen AD, Weber KL, other investigators. CRISPR-engineered T cells in patients with refractory cancer. Science, 2020; 367(6481): eaba7365.
- Yu AM, Choi YC, Tu MJ. RNA drugs and CRISPR/Cas9 therapy: Keys for tackling cancer and other diseases. Frontiers in Pharmacology, 2019; 10: 125.

