Birth Trauma in a Patient with Prader-Willi Syndrome: A Diagnostic Dilemma
Kevin Y Huang1, Pratik Gongloor1, Hai Wu2, Megan P Morand3, Joseph W Ray3 and Jianli Dong2,*
1School of Medicine, University of Texas Medical Branch, Galveston, USA
2Department of Pathology, University of Texas Medical Branch, Galveston, USA
3Department of Pediatrics, University of Texas Medical Branch, Galveston, USA
Received Date: 03/03/2025; Published Date: 28/07/2025
*Corresponding author: Jianli Dong, Department of Pathology, University of Texas Medical Branch, Galveston, TX, USA
Abstract
Prader-Willi Syndrome (PWS)—a disorder caused by the loss of paternally derived gene expression on chromosome 15—shares clinical features with numerous congenital myopathies. In neonates with PWS, elevated Creatine Kinase (CK) levels can further confound the diagnosis, particularly when perinatal trauma contributes to this laboratory finding. We now present a neonate with hypotonia and elevated CK, the combination of which raised suspicion for a primary muscular disorder instead of PWS. However, Chromosomal Microarray Analysis (CMA) and Methylation-Specific Multiplex Ligation-dependent Probe Amplification (MS-MLPA) were used to clarify the genetic basis of the disease, ultimately confirming PWS by unveiling the methylation pattern in the Prader-Willi critical region. This case emphasizes the need to consider birth trauma as a source of elevated CK in a hypotonic infant, the misinterpretation of which can delay an accurate diagnosis. Here, the use of CMA and MS-MLPA proved critical in differentiating PWS from overlapping muscular disorders, underscoring the value of these tools in cases complicated by atypical findings.
Keywords: Prader-Willi syndrome; Uniparental disomy; UPD; Chromosome microarray assay; CMA; DNA copy number variant; Methylation-specific multiplex ligation-dependent probe amplification; MS-MLPA
Introduction
Prader–Willi Syndrome (PWS) is a complex, inherited disorder with a prevalence of roughly 1 in 20,000–30,000 births globally, described first by Andrea Prader, Heinrich Willi, and Alexis Labhart in 1956 [1]. Owing to the loss of expression of paternally inherited genes in the 15q11-13 region on chromosome 15, PWS manifests with hypotonia and various other physical symptoms at birth, eventually progressing to include findings as diverse as hyperphagia, obesity, short stature, hypogonadism, intellectual delay, attention deficits, and compulsive behavioral patterns, among other clinical features [1].
Mechanistically, the loss of expression of pertinent 15q11-13 genes in PWS is associated primarily with de novo gene mutations (e.g., deletions, maternal uniparental disomy, de novo imprinting mutations) with a recurrence risk of 0–1%, although inherited imprinting defects and various other 15q chromosomal anomalies with recurrence risks of up to 50% have more rarely been identified as causative etiologies [1,2]. Importantly, molecular genetic testing is required not only for the diagnosis of PWS upon clinical suspicion at birth but also the assessment of the recurrence risk, which will be mechanism dependent [1]. Favorable near- and long-term outcomes in PWS hinge on early detection and management via tailored, multidisciplinary interventions, including hormone therapy, nutritional oversight, and behavior management [3].
Notably, elevated serum creatine kinase levels at birth, characteristic of congenital muscular diseases but not PWS, can also result from birth trauma, thereby potentially obscuring the diagnosis of PWS [4-7]. Here, we present a case of presumed birth trauma and PWS, highlighting the role of genetic testing in resolving a complex differential diagnosis.
Patient History
A 3145 g, full-term female was born at 39 weeks of gestation, pregnancy uneventful, via low transverse cesarean section. From the outset, she did not tolerate oral feeding due to oral motor dysfunction. The neonatal physical examination revealed marked hypotonia, and subsequent laboratory studies showed an elevated serum creatine kinase of 798 U/L (N: 68–580 U/L) [8]. These findings raised suspicion of a range of congenital muscular dystrophies (e.g., myotonic dystrophy, metabolic myopathies (e.g., Pompe disease), and spinal muscular atrophy. Due to its high carrier frequency in the general population, spinal muscular atrophy was a leading consideration in the differential diagnosis, and genetic testing was promptly ordered. Neither SMN1 nor SMN2, genes associated with spinal muscular atrophy, showed any pathogenic variants (tested using Invitae Spinal Muscular Atrophy STAT panel), thereby prompting a multigene neuromuscular disorder panel (tested using 211 gene Invitae Comprehensive Neuromuscular Disorder Panel) to screen for other congenital muscular disorders via sequence analysis and deletion/duplication testing. Out of the 211 genes that were evaluated, only one variant of uncertain significance, a point mutation in the PLEC gene on chromosome 8, was identified, though it is unlikely that this gene played any role in the development of the patient’s muscular abnormalities. Upon ruling out the most common causes of elevated CK and hypotonia, including myotonic dystrophy, Pompe disease (GAA), and other congenital muscular syndromes, a chromosome microarray study was performed, revealing a single copy loss of 15q11.2q13.1, including the critical regions associated with PWS or AS (Figure 1). Further testing via methylation-sensitive PCR confirmed the diagnosis of PWS specifically via detecting an abnormal paternal methylation pattern and intact maternally contributed PWS:AS critical region.
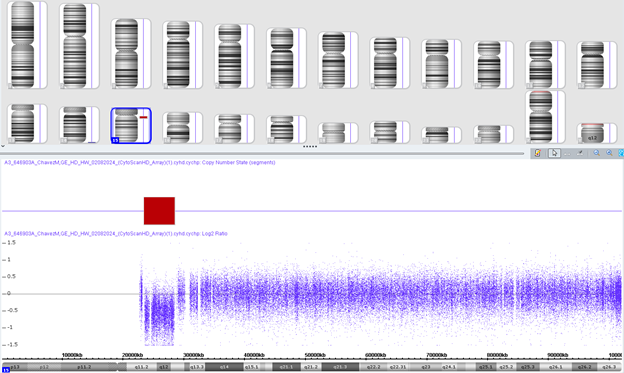
Figure 1: Chromosome microarray analysis reveals a heterozygous deletion at 15q11.2q13.1.
Top: Karyoview; Bottom: Log2 Ratio. Chromosome 15 and the deleted region are highlighted. The analysis identified a 5,040 kbp loss at 15q11.2q13.1: arr[GRCh38] 15q11.2q13.1(23375007_28414892)x1. This region encompasses multiple genes, including GOLGA6L2, MIR4508, MKRN3, MAGEL2, NDN, PWRN4, PWRN2, PWRN3, PWRN1, NPAP1, SNHG14, SNRPN, SNURF, SNORD107, PWARSN, PWAR5, SNORD64, SNORD108, PWAR6, SNORD109A, SNORD116-1 to SNORD116-30, IPW, PWAR1, SNORD115-1 to SNORD115-48 , PWAR4, SNORD109B, UBE3A, ATP10A, MIR4715, ATP10A-DT, LINC02346, LINC00929, GABRB3, GABRA5, GABRG3, GABRG3-AS1, OCA2, HERC2, GOLGA6L24, and GOLGA8F. The paternal and maternal expressed genes are indicated in blue and red, respectively.
Discussion
In this report, we present the genetic findings pertaining to a 2-month-old female patient diagnosed with PWS via Chromosome Microarray Analysis (CMA) and methylation-specific Multiplex Ligation Probe Amplification (MLPA). CMA revealed a 5,040 kbp single copy deletion of 15q11.2q13.1 on chromosome 15, consistent with PWS or AS. Further methylation-specific MLPA confirmed the 15q deletion via copy number analysis and revealed an abnormal paternal methylation pattern and intact maternally contributed PWS:AS critical region. AS was excluded based on the confirmation of the loss of the paternal allele, and the patient was diagnosed with PWS secondary to a presumed de novo deletion in the Prader-Willi Critical Region (PWCR), supported by a comprehensive family history.
CMA is considered a first-line genetic study in identifying potential chromosomal deletions and duplications spanning the entire genome, including deletions in the 15q11-q13 region specific to PWS and AS [9]. Outside the PWCR, CMA can detect other pathogenic copy number variants (CNVs), thereby potentially aiding in the diagnosis of coexisting genetic abnormalities that may influence phenotype [9]. Alternatively, Fluorescence in Situ Hybridization (FISH) enables the direct visualization of chromosomal abnormalities via fluorescently labeled DNA probes [10]. While CMA and FISH can effectively detect deletions typical of PWS, cases of PWS arising from maternal Uniparental Disomy (UPD) or imprinting defects do not involve structural CNVs and therefore evade diagnosis via these methods. Importantly, CMA and FISH are unable to distinguish between PWS and AS in most cases without further testing of parental samples [1].
Methylation-specific MLPA is a definitive diagnostic tool for PWS and AS, as well as for detecting cases of UDP and other non-CNV etiologies [11]. Aberrant methylation patterns characteristic of PWS or AS are identified via this method, allowing for a sensitive and specific analysis [12]. The absence of paternal gene expression (i.e., contribution) at the PWCR is diagnostic for PWS, encompassing the key genetic mechanisms of the disease, including deletions, UPD, and imprinting defects [1].
Deletion breakpoints in PWS are used to subclassify the disease and aid in genotype–phenotype correlations [2]. Type 1 and Type 2 deletions share the distal breakpoint BP3 and break off proximally at BP1 and BP2, respectively [1]. The patient here was found to have a loss of MKRN3 distal to BP1 and HERC2 proximal to BP2, with no further loss of gene outside of the BP2–BP3 region, thus qualifying the deletion as Type 2. Phenotypically, the patient lacked certain characteristic facial features of PWS, such as narrowing of the temples, strabismus, and a thin upper lip [13]. A different case of PWS involving the BP2–BP3 region, as well as the more distal APBA2 and CHRNA7 genes near BP4 and BP5, was also characterized by a lack of the typical facial features of PWS, though importantly, macrocephaly was noted, even prior to initiating growth hormone therapy [2]. Additionally, frontal bossing and large hands and feet were observed, perhaps suggesting that certain pertinent genes upstream of BP3 are responsible for the discrepancy in dysmorphic features [2]. Proximal to BP2, the non-imprinted genes NIPA1, NIPA2, and CYFIP1 were spared in the present, normocephalic case of PWS. At least two reported cases of PWS involving the loss of BP1–BP2 have noted the atypical finding of microcephaly [1,2], thereby implicating the loss of NIPA1, NIPA2, and CYFIP1, or some combination thereof, in the development of this particular clinical feature. Further laboratory investigations into the genetic mechanisms driving the spectrum of PWS presentation should reveal clearer insights into the genotype–phenotype relationship.
This case highlights an unusual presentation of PWS where the initial presentation of hypotonia and markedly elevated CK could have led the physician astray if not for thorough genetic testing and diagnostic acumen. Importantly, PWS is not typically associated with CK levels above the normal range for healthy newborns, though one rare case of elevated CK of unknown origin has been reported [5,14]. In the absence of any congenital muscular disorder, it is well established that the elevated CK could be attributed to perinatal trauma arising from the elective cesarean section [15]. It has been suggested that a serum CK in the range of 805.8 ± 465.2 U/L is consistent with such a scenario [6]. The identification of atypical presentations of PWS with concomitant birth trauma is critical for early diagnosis and intervention, particularly with respect to ruling out adjacent neuromuscular disorders.
In this case, the abnormal creatine kinase level adds a unique dimension to the spectrum of PWS presentations, underscoring the role of laboratory investigation in resolving a complex neurodevelopmental and neuromuscular differential diagnosis.
References
- Driscoll DJ, Miller JL, Cassidy SB. Prader-Willi syndrome. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, eds. GeneReviews®. University of Washington, Seattle, 2024.
- Kim SJ, Miller JL, Kuipers PJ, et al. Unique and atypical deletions in Prader–Willi syndrome reveal distinct phenotypes. Eur J Hum Genet, 2012; 20(3): 283-290. doi:10.1038/ejhg.2011.187.
- Tauber M, Hoybye C. Endocrine disorders in Prader-Willi syndrome: a model to understand and treat hypothalamic dysfunction. The Lancet Diabetes & Endocrinology, 2021; 9(4): 235-246. doi:10.1016/S2213-8587(21)00002-4.
- Bodensteiner JB. The Evaluation of the Hypotonic Infant. Seminars in Pediatric Neurology, 2008; 15(1): 10-20. doi: 10.1016/j.spen.2008.01.003.
- Al-Ghamdi F, Ghosh PS. Clinical Reasoning: A tale of a hypotonic infant. Neurology, 2016; 87(2). doi: 10.1212/WNL.0000000000002833
- Al-twaijri WA, Al-saif SA, Al-fehaid GI, Al-qirnas MQ, Masuadi EM. Elevated level of creatine phosphokinase in newborn: Clinical significance and association with congenital muscle diseases. NSJ, 2022; 27(4): 263-269. doi: 10.17712/nsj.2022.4.20220051.
- Maloney B, Park S, Sowizral M, et al. Factors influencing creatine kinase-MM concentrations in newborns and implications for newborn screening for Duchenne muscular dystrophy. Clinical Biochemistry, 2023; 118: 110614. doi: 10.1016/j.clinbiochem.2023.110614.
- Pagana KD. Mosby’s® Diagnostic and Laboratory Test Reference - E-Book. 2020; 15th ed. Mosby.
- Dugoff L, Norton ME, Kuller JA. The use of chromosomal microarray for prenatal diagnosis. American Journal of Obstetrics and Gynecology, 2016; 215(4): B2-B9. doi: 10.1016/j.ajog.2016.07.016.
- Cui C, Shu W, Li P. Fluorescence In situ Hybridization: Cell-Based Genetic Diagnostic and Research Applications. Front Cell Dev Biol, 2016; 4. doi: 10.3389/fcell.2016.00089.
- Nygren AOH. Methylation-Specific MLPA (MS-MLPA): simultaneous detection of CpG methylation and copy number changes of up to 40 sequences. Nucleic Acids Research, 2005; 33(14): e128-e128. doi: 10.1093/nar/gni127.
- Henkhaus RS, Kim SJ, Kimonis VE, et al. Methylation-Specific Multiplex Ligation-Dependent Probe Amplification and Identification of Deletion Genetic Subtypes in Prader-Willi Syndrome. Genetic Testing and Molecular Biomarkers, 2012; 16(3): 178-186. doi: 10.1089/gtmb.2011.0115.
- Aughton DJ. Physical Features of Prader-Willi Syndrome in Neonates. Arch Pediatr Adolesc Med, 1990; 144(11): 1251. doi: 10.1001/archpedi.1990.02150350083032.
- McCandless SE. Prader-Willi Syndrome. NeoReviews, 2005; 6(12): e559-e566. doi: 10.1542/neo.6-12-e559.
- Rudolph N, Gross RT. Creatine phosphokinase activity in serum of newborn infants as an indicator of fetal trauma during birth. Pediatrics, 1966; 38(6): 1039-1046. doi: 10.1542/peds.38.6.1039.

